Standardization¶
This page gives details on the standardization process.
Standardizing a molecule¶
The standardize_smiles function provides a quick and easy way to get the standardized version of a given SMILES
string:
>>> from molvs import standardize_smiles
>>> standardize_smiles('C[n+]1c([N-](C))cccc1')
'CN=c1ccccn1C'
While this is convenient for one-off cases, it’s inefficient when dealing with multiple molecules and doesn’t allow any customization of the standardization process.
The Standardizer class provides flexibility to specify custom standardization stages and efficiently standardize
multiple molecules:
>>> from rdkit import Chem
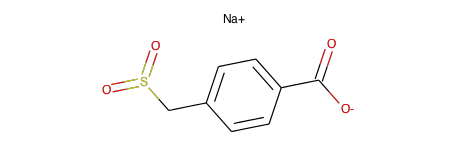
>>> mol = Chem.MolFromSmiles('[Na]OC(=O)c1ccc(C[S+2]([O-])([O-]))cc1')

>>> from molvs import Standardizer
>>> s = Standardizer()
>>> smol = s.standardize(mol)
The standardization process¶
TODO: Explain this properly…
RDKit Sanitize¶
- Nitro N=O:
CN(=O)=O >> C[N+](=O)[O-]andC1=CC=CN(=O)=C1 >> C1=CC=C[N+]([O-])=C1 - Nitro N#O:
C-N=N#N >> C-N=[N+]=[N-] - Perchlorate:
Cl(=O)(=O)(=O)[O-] >> [Cl+3]([O-])([O-])([O-])[O-] - Calculate explicit and implicit valence of all atoms. Fails when atoms have illegal valence.
- Calculate symmetrized SSSR. Slowest step, fails in rare cases.
- Kekulize. Fails if a Kekule form cannot be found or non-ring bonds are marked as aromatic.
- Assign radicals if hydrogens set and bonds+hydrogens+charge < valence.
- Set aromaticity, if none set in input. Go round rings, Huckel rule to set atoms+bonds as aromatic.
- Set conjugated property on bonds where applicable.
- Set hybridisation property on atoms.
- Remove chirality markers from sp and sp2 hybridised centers.
RDKit RemoveHs¶
- RDKit implementation detail - this is the preferred way to store the molecule.
- Remove explicit H count from atoms, instead infer it on the fly from valence model.
Disconnect metals¶
- Break covalent bonds between metals and organic atoms under certain conditions.
- First, disconnect N, O, F from any metal. Then disconnect other non-metals from transition metals (with exceptions).
- For every bond broken, adjust the charges of the begin and end atoms accordingly.
- In future, we might attempt to replace with zero-order bonds.
Apply normalization rules¶
- A series of transformations to correct common drawing errors and standardize functional groups. Includes:
- Uncharge-separate sulfones
- Charge-separate nitro groups
- Charge-separate pyridine oxide
- Charge-separate azide
- Charge-separate diazo and azo groups
- Charge-separate sulfoxides
- Hydrazine-diazonium system
Reionize acids¶
If molecule with multiple acid groups is partially ionized, ensure strongest acids ionize first.
The algorithm works as follows:
- Use SMARTS to find the strongest protonated acid and the weakest ionized acid.
- If the ionized acid is weaker than the protonated acid, swap proton and repeat.
Recalculate stereochemistry¶
- Use built-in RDKit functionality to force a clean recalculation of stereochemistry